Exploring Functional Brain Networks in Alzheimer’s Disease Using Resting State EEG Signals
Abstract
1. Introduction
- The examination of the usefulness of resting EEG signals, under the functional connectivity concept, to discriminate between HC and SCD using a large cohort of subjects (∼1000).
- The evaluation of connectivity metrics over the entire AD spectrum, including people from the SCD group, by performing statistical analysis and discriminant analysis.
- In most works, few graph metrics were reported in common, precluding meta-analyses. In our analysis, we include a large number of metrics; hence, we provide a holistic view of our study with respect to brain’s connectivity.
- Finally, we provide a holistic view of data analysis by incorporating statistical analysis, topographical visualization, and predictive analysis.
2. Materials and Methods
2.1. Subjects
2.2. EEG Recordings
2.3. General Procedure for Network Construction
2.3.1. Connectivity as a General Measure
2.3.2. Constructing the Adjacency Matrix
2.3.3. Thresholding the Matrix
2.4. Connectivity Measures
2.4.1. Pearson Correlation (CORR)
- and are vectors representing the EEG signals (/time—series) from channels and .
- and are the mean values of the signals and .
2.4.2. Phase Locking Value (PLV)
- and are vectors representing the EEG signals from channels and .
- and are the instantaneous phases of and at time .
2.5. Connectivity (Graph Theory) Metrics
- represents the number of connections between neighbors of node .
- is the degree of node (number of neighbors).
- is the adjacency matrix element representing the connection between nodes and .
- is the largest eigenvalue of .
- is the eigenvector centrality of node .
- is the weight of the edge between nodes and .
- and : Strengths of nodes and , calculated as , where is the weight of the edge between and .
- : Fraction of total edge weight that the connection between and contributes, computed as .
- is the total number of shortest paths between nodes and .
- is the number of those paths that pass through node .
- Note: The sum is taken over all node pairs where and .
- is the total number of nodes in the network.
- is the shortest path distance between nodes and .
2.6. Analysis of EEG Time Series
2.6.1. Group-Level Comparison with ANCOVA for Controlling Age Effects
2.6.2. Topographical Visualization of Group-Specific Brain Network Patterns
2.6.3. Machine Learning Tasks for the Discrimination of AD Stages Based on Connectivity Metrics
3. Experiments and Results
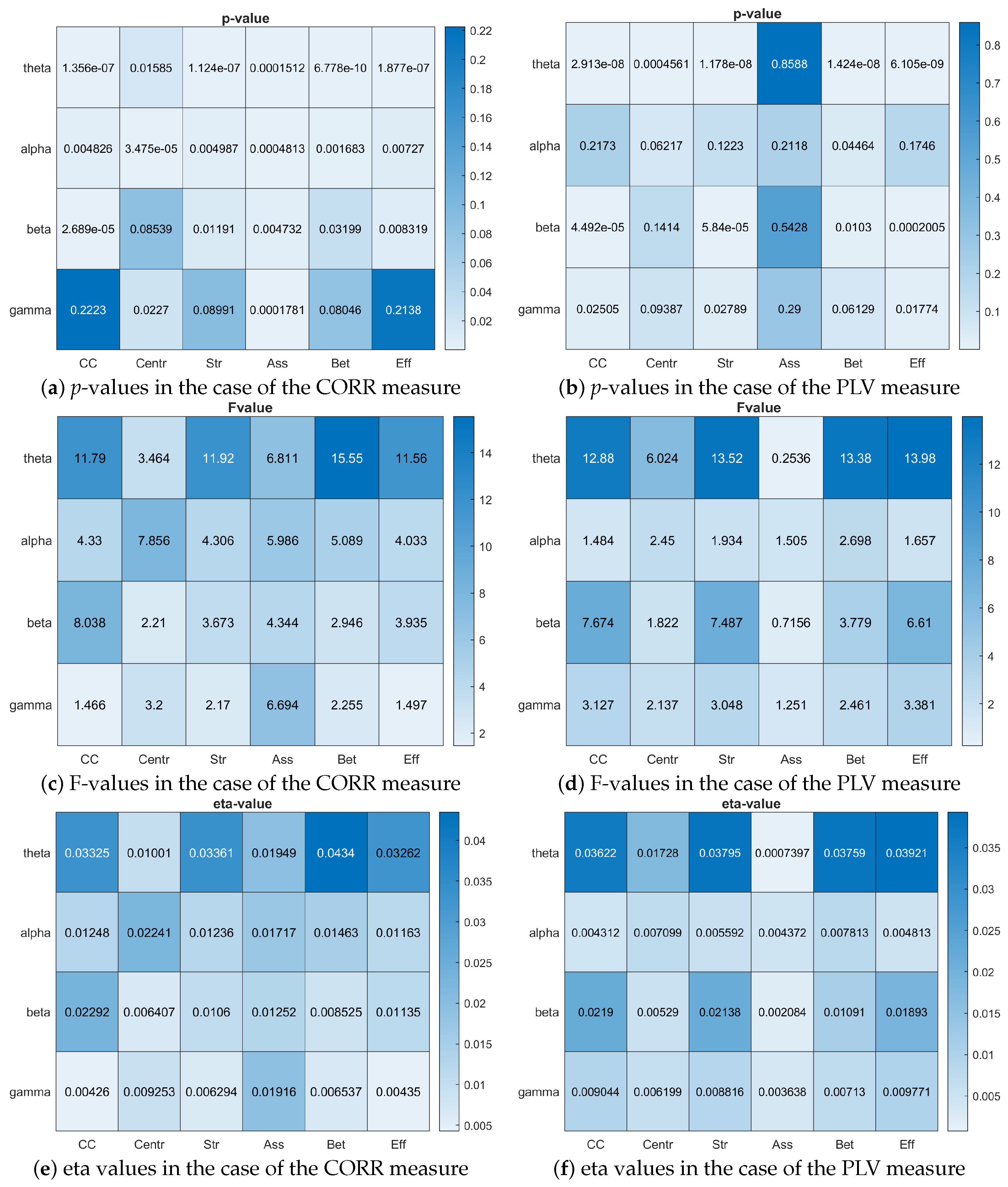
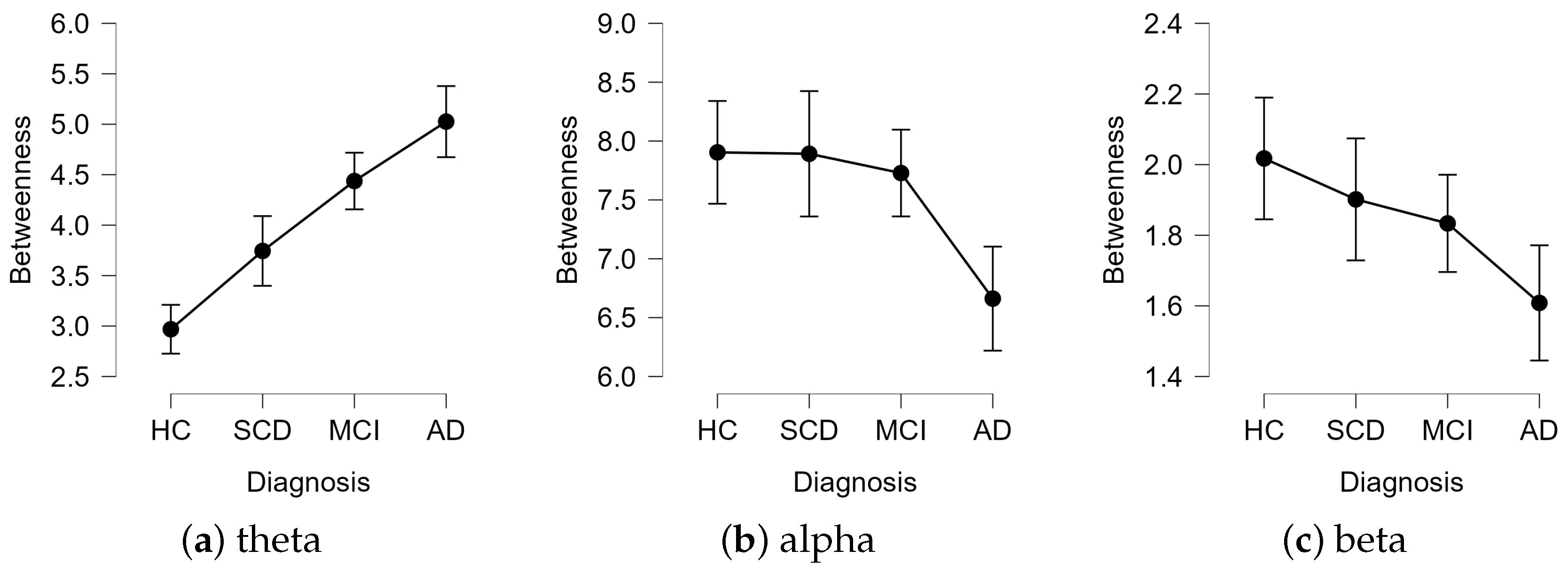
3.1. Group-Level Comparison
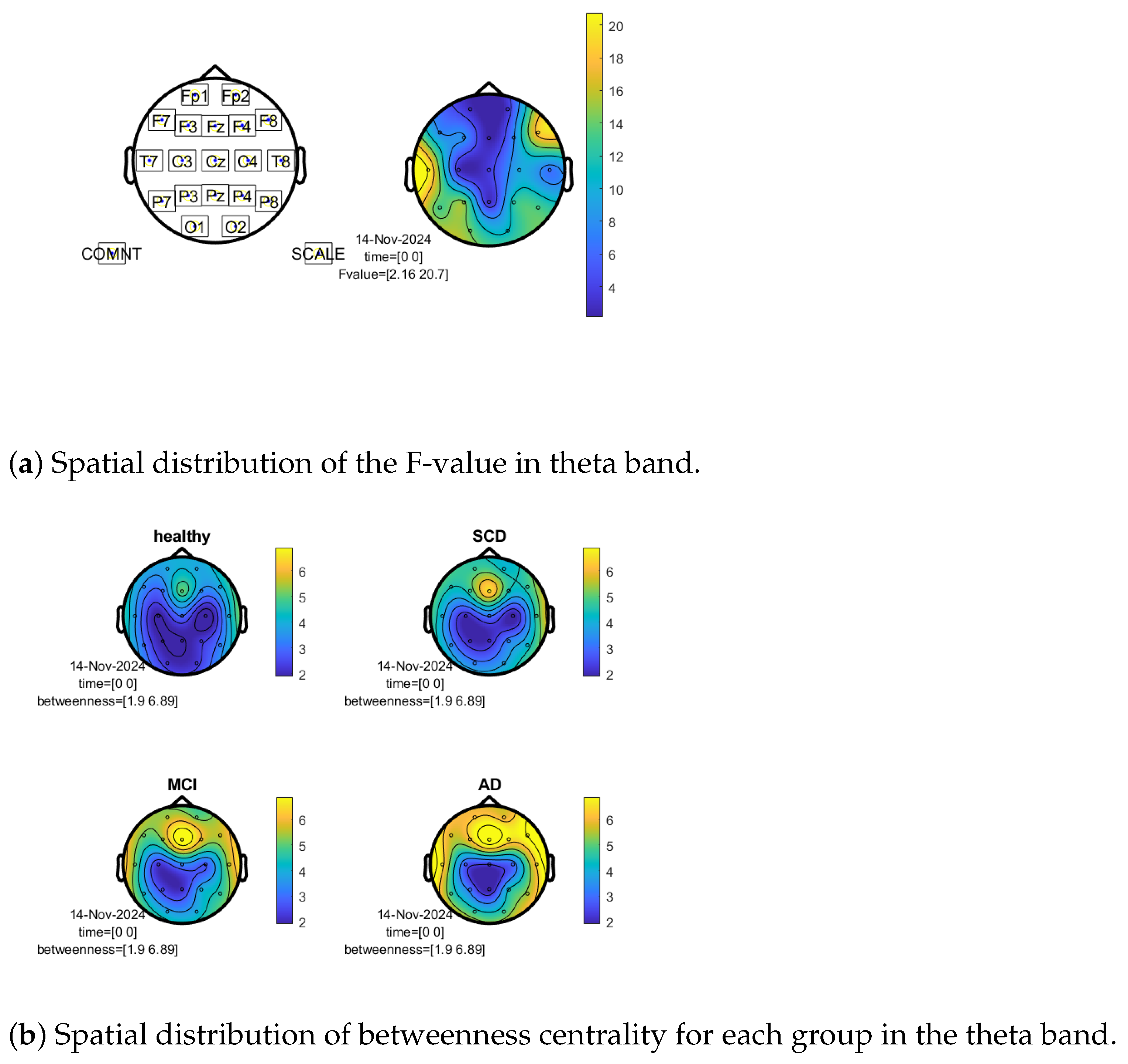
3.2. Topographical Visualization
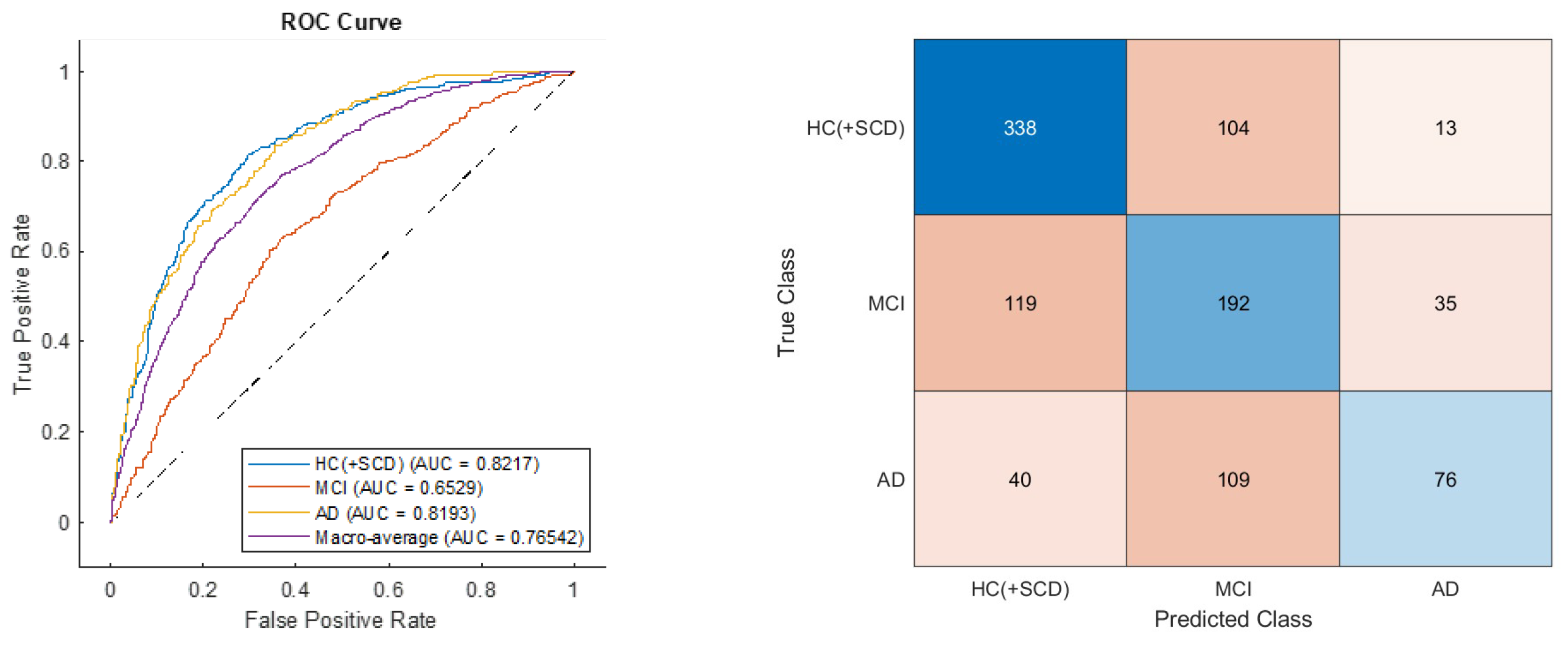
3.3. Classification of AD Stages Based on Connectivity Metrics
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Correction Statement
References
- Jack, C.R.; Bennett, D.A.; Blennow, K.; Carrillo, M.C.; Dunn, B.; Haeberlein, S.B.; Holtzman, D.M.; Jagust, W.; Jessen, F.; Karlawish, J.; et al. NIA-AA Research Framework: Toward a biological definition of Alzheimer’s disease. Alzheimer’s Dement. 2018, 14, 535–562. [Google Scholar] [CrossRef] [PubMed]
- Alberdi, A.; Aztiria, A.; Basarab, A. On the early diagnosis of Alzheimer’s Disease from multimodal signals: A survey. Artif. Intell. Med. 2016, 71, 1–29. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.K.; Kim, H.; Kim, S.H.; Kim, J.B.; Kim, L. Electroencephalography-based classification of Alzheimer’s disease spectrum during computer-based cognitive testing. Sci. Rep. 2024, 14, 5252. [Google Scholar] [CrossRef]
- Jessen, F.; Amariglio, R.E.; Van Boxtel, M.; Breteler, M.; Ceccaldi, M.; Chételat, G.; Dubois, B.; Dufouil, C.; Ellis, K.A.; Van Der Flier, W.M.; et al. A conceptual framework for research on subjective cognitive decline in preclinical Alzheimer’s disease. Alzheimer’s Dement. 2014, 10, 844–852. [Google Scholar] [CrossRef]
- Sperling, R.; Mormino, E.; Johnson, K. The Evolution of Preclinical Alzheimer’s Disease: Implications for Prevention Trials. Neuron 2014, 84, 608–622. [Google Scholar] [CrossRef]
- Jessen, F.; Amariglio, R.E.; Buckley, R.F.; Van Der Flier, W.M.; Han, Y.; Molinuevo, J.L.; Rabin, L.; Rentz, D.M.; Rodriguez-Gomez, O.; Saykin, A.J.; et al. The characterisation of subjective cognitive decline. Lancet Neurol. 2020, 19, 271–278. [Google Scholar] [CrossRef] [PubMed]
- Si, T.; Xing, G.; Han, Y. Subjective Cognitive Decline and Related Cognitive Deficits. Front. Neurol. 2020, 11, 247. [Google Scholar] [CrossRef]
- Cassani, R.; Estarellas, M.; San-Martin, R.; Fraga, F.J.; Falk, T.H. Systematic Review on Resting-State EEG for Alzheimer’s Disease Diagnosis and Progression Assessment. Dis. Markers 2018, 2018, 5174815. [Google Scholar] [CrossRef]
- Zheng, X.; Wang, B.; Liu, H.; Wu, W.; Sun, J.; Fang, W.; Jiang, R.; Hu, Y.; Jin, C.; Wei, X.; et al. Diagnosis of Alzheimer’s disease via resting-state EEG: Integration of spectrum, complexity, and synchronization signal features. Front. Aging Neurosci. 2023, 15, 1288295. [Google Scholar] [CrossRef]
- Sharma, N.; Kolekar, M.; Jha, K.; Kumar, Y. EEG and Cognitive Biomarkers Based Mild Cognitive Impairment Diagnosis. IRBM 2019, 40, 113–121. [Google Scholar] [CrossRef]
- Abazid, M.; Houmani, N.; Boudy, J.; Dorizzi, B.; Mariani, J.; Kinugawa, K. A Comparative Study of Functional Connectivity Measures for Brain Network Analysis in the Context of AD Detection with EEG. Entropy 2021, 23, 1553. [Google Scholar] [CrossRef] [PubMed]
- Şeker, M.; Özbek, Y.; Yener, G.; Özerdem, M.S. Complexity of EEG Dynamics for Early Diagnosis of Alzheimer’s Disease Using Permutation Entropy Neuromarker. Comput. Methods Programs Biomed. 2021, 206, 106116. [Google Scholar] [CrossRef] [PubMed]
- Su, R.; Li, X.; Li, Z.; Han, Y.; Cui, W.; Xie, P.; Liu, Y. Constructing biomarker for early diagnosis of aMCI based on combination of multiscale fuzzy entropy and functional brain connectivity. Biomed. Signal Process. Control 2021, 70, 103000. [Google Scholar] [CrossRef]
- Stam, C.J. Modern network science of neurological disorders. Nat. Rev. Neurosci. 2014, 15, 683–695. [Google Scholar] [CrossRef]
- Vecchio, F.; Miraglia, F.; Maria Rossini, P. Connectome: Graph theory application in functional brain network architecture. Clin. Neurophysiol. Pract. 2017, 2, 206–213. [Google Scholar] [CrossRef] [PubMed]
- Friston, K.J. Functional and Effective Connectivity: A Review. Brain Connect. 2011, 1, 13–36. [Google Scholar] [CrossRef]
- Stam, C.J. Hub overload and failure as a final common pathway in neurological brain network disorders. Netw. Neurosci. 2024, 8, 1–23. [Google Scholar] [CrossRef]
- Rubinov, M.; Sporns, O. Complex network measures of brain connectivity: Uses and interpretations. NeuroImage 2010, 52, 1059–1069. [Google Scholar] [CrossRef]
- Vecchio, F.; Miraglia, F.; Quaranta, D.; Granata, G.; Romanello, R.; Marra, C.; Bramanti, P.; Rossini, P. Cortical connectivity and memory performance in cognitive decline: A study via graph theory from EEG data. Neuroscience 2016, 316, 143–150. [Google Scholar] [CrossRef]
- Chetty, C.A.; Bhardwaj, H.; Kumar, G.P.; Devanand, T.; Sekhar, C.S.A.; Aktürk, T.; Kiyi, I.; Yener, G.; Güntekin, B.; Joseph, J.; et al. EEG biomarkers in Alzheimer’s and prodromal Alzheimer’s: A comprehensive analysis of spectral and connectivity features. Alzheimer’s Res. Ther. 2024, 16, 236. [Google Scholar] [CrossRef]
- Vecchio, F.; Miraglia, F.; Marra, C.; Quaranta, D.; Vita, M.G.; Bramanti, P.; Rossini, P.M. Human Brain Networks in Cognitive Decline: A Graph Theoretical Analysis of Cortical Connectivity from EEG Data. J. Alzheimer’s Dis. 2014, 41, 113–127. [Google Scholar] [CrossRef] [PubMed]
- Miraglia, F.; Vecchio, F.; Bramanti, P.; Rossini, P.M. EEG characteristics in “eyes-open” versus “eyes-closed” conditions: Small-world network architecture in healthy aging and age-related brain degeneration. Clin. Neurophysiol. 2016, 127, 1261–1268. [Google Scholar] [CrossRef]
- Fodor, Z.; Horváth, A.; Hidasi, Z.; Gouw, A.A.; Stam, C.J.; Csukly, G. EEG Alpha and Beta Band Functional Connectivity and Network Structure Mark Hub Overload in Mild Cognitive Impairment During Memory Maintenance. Front. Aging Neurosci. 2021, 13, 680200. [Google Scholar] [CrossRef] [PubMed]
- Cai, L.; Wei, X.; Liu, J.; Zhu, L.; Wang, J.; Deng, B.; Yu, H.; Wang, R. Functional Integration and Segregation in Multiplex Brain Networks for Alzheimer’s Disease. Front. Neurosci. 2020, 14, 51. [Google Scholar] [CrossRef]
- Das, S.; Puthankattil, S.D. Complex network analysis of MCI-AD EEG signals under cognitive and resting state. Brain Res. 2020, 1735, 146743. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Zhao, J.; Chen, X.; Yin, B.; Li, X.; Xie, P. Alzheimer’s disease diagnosis using rhythmic power changes and phase differences: A low-density EEG study. Front. Aging Neurosci. 2025, 16, 1485132. [Google Scholar] [CrossRef]
- Kim, M.J.; Youn, Y.C.; Paik, J. Deep learning-based EEG analysis to classify normal, mild cognitive impairment, and dementia: Algorithms and dataset. NeuroImage 2023, 272, 120054. [Google Scholar] [CrossRef] [PubMed]
- Hallquist, M.N.; Hillary, F.G. Graph theory approaches to functional network organization in brain disorders: A critique for a brave new small-world. Netw. Neurosci. 2019, 3, 1–26. [Google Scholar] [CrossRef]
- Tang, H.; Ma, G.; Zhang, Y.; Ye, K.; Guo, L.; Liu, G.; Huang, Q.; Wang, Y.; Ajilore, O.; Leow, A.D.; et al. A comprehensive survey of complex brain network representation. Meta-Radiology 2023, 1, 100046. [Google Scholar] [CrossRef]
- Farahani, F.V.; Karwowski, W.; Lighthall, N.R. Application of Graph Theory for Identifying Connectivity Patterns in Human Brain Networks: A Systematic Review. Front. Neurosci. 2019, 13, 585. [Google Scholar] [CrossRef]
- Adamovich, T.; Zakharov, I.; Tabueva, A.; Malykh, S. The thresholding problem and variability in the EEG graph network parameters. Sci. Rep. 2022, 12, 18659. [Google Scholar] [CrossRef]
- Sporns, O. Graph theory methods: Applications in brain networks. Dialogues Clin. Neurosci. 2018, 20, 111–121. [Google Scholar] [CrossRef] [PubMed]
- Joyce, K.E.; Laurienti, P.J.; Burdette, J.H.; Hayasaka, S. A New Measure of Centrality for Brain Networks. PLoS ONE 2010, 5, e12200. [Google Scholar] [CrossRef] [PubMed]
- Blum, S.; Jacobsen, N.S.J.; Bleichner, M.G.; Debener, S. A Riemannian Modification of Artifact Subspace Reconstruction for EEG Artifact Handling. Front. Hum. Neurosci. 2019, 13, 141. [Google Scholar] [CrossRef] [PubMed]
- Dauwels, J.; Vialatte, F.B.; Cichocki, A. On the Early Diagnosis of Alzheimer’s Disease from EEG Signals: A Mini-Review. In Advances in Cognitive Neurodynamics (II); Wang, R., Gu, F., Eds.; Springer: Dordrecht, The Netherlands, 2011; pp. 709–716. [Google Scholar] [CrossRef]
- Yang, X.; Fan, Z.; Li, Z.; Zhou, J. Resting-state EEG microstate features for Alzheimer’s disease classification. PLoS ONE 2024, 19, e0311958. [Google Scholar] [CrossRef]
- Tan, E.; Troller-Renfree, S.V.; Morales, S.; Buzzell, G.A.; McSweeney, M.; Antúnez, M.; Fox, N.A. Theta activity and cognitive functioning: Integrating evidence from resting-state and task-related developmental electroencephalography (EEG) research. Dev. Cogn. Neurosci. 2024, 67, 101404. [Google Scholar] [CrossRef]
- Oikonomou, V.P.; Geordiadis, K.; Kalaganis, F.P.; Nikolopoulos, S.; Kompatsiaris, I. Prediction of Successful Memory Formation during Audiovisual advertising using EEG signals. In Proceedings of the 2024 IEEE Conference on Artificial Intelligence (CAI), Singapore, 25–27 June 2024; pp. 1111–1116. [Google Scholar] [CrossRef]
- Weiss, S.; Mueller, H.M. “Too Many betas do not Spoil the Broth”: The Role of Beta Brain Oscillations in Language Processing. Front. Psychol. 2012, 3, 201. [Google Scholar] [CrossRef]
- Foxe, J.J.; Snyder, A.C. The Role of Alpha-Band Brain Oscillations as a Sensory Suppression Mechanism during Selective Attention. Front. Psychol. 2011, 2, 154. [Google Scholar] [CrossRef] [PubMed]
- Hallett, M.; De Haan, W.; Deco, G.; Dengler, R.; Di Iorio, R.; Gallea, C.; Gerloff, C.; Grefkes, C.; Helmich, R.C.; Kringelbach, M.L.; et al. Human brain connectivity: Clinical applications for clinical neurophysiology. Clin. Neurophysiol. 2020, 131, 1621–1651. [Google Scholar] [CrossRef]
- Sun, Y.; Yin, Q.; Fang, R.; Yan, X.; Wang, Y.; Bezerianos, A.; Tang, H.; Miao, F.; Sun, J. Disrupted Functional Brain Connectivity and Its Association to Structural Connectivity in Amnestic Mild Cognitive Impairment and Alzheimer’s Disease. PLoS ONE 2014, 9, e96505. [Google Scholar] [CrossRef]
- Lazarou, I.; Georgiadis, K.; Nikolopoulos, S.; Oikonomou, V.P.; Tsolaki, A.; Kompatsiaris, I.; Tsolaki, M.; Kugiumtzis, D. A Novel Connectome-based Electrophysiological Study of Subjective Cognitive Decline Related to Alzheimer’s Disease by Using Resting-state High-density EEG EGI GES 300. Brain Sci. 2020, 10, 392. [Google Scholar] [CrossRef]
- Phillips, D.J.; McGlaughlin, A.; Ruth, D.; Jager, L.R.; Soldan, A. Graph theoretic analysis of structural connectivity across the spectrum of Alzheimer’s disease: The importance of graph creation methods. Neuroimage Clin. 2015, 7, 377–390. [Google Scholar] [CrossRef] [PubMed]
- Frantzidis, C.A.; Vivas, A.B.; Tsolaki, A.; Klados, M.A.; Tsolaki, M.; Bamidis, P.D. Functional disorganization of small-world brain networks in mild Alzheimer’s Disease and amnestic Mild Cognitive Impairment: An EEG study using Relative Wavelet Entropy (RWE). Front. Aging Neurosci. 2014, 6, 224. [Google Scholar] [CrossRef]
- RADAR-AD Consortium; Lazarou, I.; Georgiadis, K.; Nikolopoulos, S.; Oikonomou, V.P.; Stavropoulos, T.G.; Tsolaki, A.; Kompatsiaris, I.; Tsolaki, M. Exploring Network Properties Across Preclinical Stages of Alzheimer’s Disease Using a Visual Short-Term Memory and Attention Task with High-Density Electroencephalography: A Brain-Connectome Neurophysiological Study. J. Alzheimer’s Dis. 2022, 87, 643–664. [Google Scholar] [CrossRef]
- Engels, M.M.; Stam, C.J.; Van Der Flier, W.M.; Scheltens, P.; De Waal, H.; Van Straaten, E.C. Declining functional connectivity and changing hub locations in Alzheimer’s disease: An EEG study. BMC Neurol. 2015, 15, 145. [Google Scholar] [CrossRef] [PubMed]
- Scheijbeler, E.P.; De Haan, W.; Stam, C.J.; Twisk, J.W.R.; Gouw, A.A. Longitudinal resting-state EEG in amyloid-positive patients along the Alzheimer’s disease continuum: Considerations for clinical trials. Alzheimer’s Res. Ther. 2023, 15, 182. [Google Scholar] [CrossRef] [PubMed]
- Mårtensson, G.; Pereira, J.B.; Mecocci, P.; Vellas, B.; Tsolaki, M.; Kłoszewska, I.; Soininen, H.; Lovestone, S.; Simmons, A.; Volpe, G.; et al. Stability of graph theoretical measures in structural brain networks in Alzheimer’s disease. Sci. Rep. 2018, 8, 11592. [Google Scholar] [CrossRef]
- Santiago, J.A.; Potashkin, J.A. The Impact of Disease Comorbidities in Alzheimer’s Disease. Front. Aging Neurosci. 2021, 13, 631770. [Google Scholar] [CrossRef]
- Oikonomou, V.P.; Kompatsiaris, I. A Novel Bayesian Approach for EEG Source Localization. Comput. Intell. Neurosci. 2020, 2020, 8837954. [Google Scholar] [CrossRef]
- Aoki, Y.; Takahashi, R.; Suzuki, Y.; Pascual-Marqui, R.D.; Kito, Y.; Hikida, S.; Maruyama, K.; Hata, M.; Ishii, R.; Iwase, M.; et al. EEG resting-state networks in Alzheimer’s disease associated with clinical symptoms. Sci. Rep. 2023, 13, 3964. [Google Scholar] [CrossRef]


| Cases | Sum of Squares | df | Mean Square | F | p | 2 |
|---|---|---|---|---|---|---|
| Diagnosis | 281.320 | 3 | 93.773 | 15.547 | <0.001 | 0.043 |
| Age | 70.211 | 1 | 70.211 | 11.641 | <0.001 | 0.011 |
| Residuals | 6200.493 | 1028 | 6.032 |
| Mean Difference | SE | t | |||
|---|---|---|---|---|---|
| HC | SCD | −0.540 | 0.244 | −2.215 | 0.120 |
| MCI | −1.124 | 0.223 | −5.032 | <0.001 *** | |
| AD | −1.636 | 0.253 | −6.474 | <0.001 *** | |
| SCD | MCI | −0.584 | 0.226 | −2.587 | 0.048 * |
| AD | −1.096 | 0.248 | −4.410 | <0.001 *** | |
| MCI | AD | −0.512 | 0.210 | −2.433 | 0.072 |
| Cases | Sum of Squares | df | Mean Square | F | p | 2 |
|---|---|---|---|---|---|---|
| Diagnosis | 192.164 | 3 | 64.055 | 5.089 | 0.002 | 0.015 |
| Age | 8.598 | 1 | 8.598 | 0.683 | 0.409 | 6.6 × 10−4 |
| Residuals | 12939.253 | 1028 | 12.587 |
| Cases | Sum of Squares | df | Mean Square | F | p | 2 |
|---|---|---|---|---|---|---|
| Diagnosis | 15.235 | 3 | 5.078 | 2.946 | 0.032 | 0.009 |
| Age | 0.179 | 1 | 0.179 | 0.104 | 0.748 | 1.0 × 10−4 |
| Residuals | 1771.756 | 1028 | 1.723 |
| Mean Difference | SE | t | |||
|---|---|---|---|---|---|
| HC | SCD | −0.070 | 0.352 | −0.020 | 0.997 |
| MCI | 0.055 | 0.323 | 0.015 | 0.998 | |
| AD | 1.095 | 0.365 | 0.309 | 0.015 * | |
| SCD | MCI | 0.125 | 0.326 | 0.035 | 0.981 |
| AD | 1.165 | 0.359 | 0.328 | 0.007 ** | |
| MCI | AD | 1.040 | 0.304 | 0.293 | 0.004 ** |
| Mean Difference | SE | t | |||
|---|---|---|---|---|---|
| HC | SCD | 0.104 | 0.130 | 0.798 | 0.855 |
| MCI | 0.166 | 0.119 | 1.394 | 0.503 | |
| AD | 0.388 | 0.135 | 2.871 | 0.022 * | |
| SCD | MCI | 0.062 | 0.121 | 0.518 | 0.955 |
| AD | 0.284 | 0.133 | 2.136 | 0.142 | |
| MCI | AD | 0.221 | 0.112 | 1.968 | 0.201 |
| Classifier | Precision | Accuracy | Sensitivity | Specificity | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| HC | MCI | AD | HC | MCI | AD | HC | MCI | AD | ||
| kNN [27] | 36.80% | 47.58% | 58.46% | 68.49% | 74.00% | 21.01% | 11.99% | 32.28% | 79.96% | 88.63% |
| SVM [27] | 52.33% | 73.47% | 59.06% | 72.14% | 77.15% | 39.44% | 32.55% | 71.11% | 69.50% | 86.25% |
| Ieracitano-CNN [27] | 54.27% | 70.86% | 61.51% | 76.17% | 76.35% | 37.14% | 44.15% | 67.35% | 74.48% | 87.58% |
| Brain Connectivity | 59.21% | 76.66% | 62.59% | 78.01% | 69.72% | 55.64% | 46.30% | 81.04% | 66.36% | 89.27% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oikonomou, V.P.; Georgiadis, K.; Lazarou, I.; Nikolopoulos, S.; Kompatsiaris, I.; PREDICTOM Consortium. Exploring Functional Brain Networks in Alzheimer’s Disease Using Resting State EEG Signals. J. Dement. Alzheimer's Dis. 2025, 2, 12. https://doi.org/10.3390/jdad2020012
Oikonomou VP, Georgiadis K, Lazarou I, Nikolopoulos S, Kompatsiaris I, PREDICTOM Consortium. Exploring Functional Brain Networks in Alzheimer’s Disease Using Resting State EEG Signals. Journal of Dementia and Alzheimer's Disease. 2025; 2(2):12. https://doi.org/10.3390/jdad2020012
Chicago/Turabian StyleOikonomou, Vangelis P., Kostas Georgiadis, Ioulietta Lazarou, Spiros Nikolopoulos, Ioannis Kompatsiaris, and PREDICTOM Consortium. 2025. "Exploring Functional Brain Networks in Alzheimer’s Disease Using Resting State EEG Signals" Journal of Dementia and Alzheimer's Disease 2, no. 2: 12. https://doi.org/10.3390/jdad2020012
APA StyleOikonomou, V. P., Georgiadis, K., Lazarou, I., Nikolopoulos, S., Kompatsiaris, I., & PREDICTOM Consortium. (2025). Exploring Functional Brain Networks in Alzheimer’s Disease Using Resting State EEG Signals. Journal of Dementia and Alzheimer's Disease, 2(2), 12. https://doi.org/10.3390/jdad2020012








